
iGEM Bonn
iGEM Bonn 2026 Projektbeschreibung

Viral protease-based plant defense tool against potyviruses
- Construction of a plant recognition and resistance tool against potyviruses
- Utilization of cleavage sites of viral proteases to measure virus abundance
- Trigger signal resulting in induce cell death response
Viral protease-based plant defense tool against potyviruses
We are currently working on a plant recognition and resistance tool against potyviruses. Plant-targeting virus-based annual global economic losses are estimated around 30 billion US dollar. (1) The potyvirus family is the largest group of viral plant pathogens (1) with most likely a significant share in that economic losses. Interestingly, the viral genome of potyviruses contains a large polyprotein, which is proteolytically processed by viral proteases into 10 mature proteins and several precursors. (1) Nla-Pro cleaves as the main protease a total of 7 cleavages sides. (3). The conservation of the Nla-protease as well as their conserved cleavage sites are features, which we want to exploit in our tool.
For screening for the virus we are utilizing a post translational modification process called
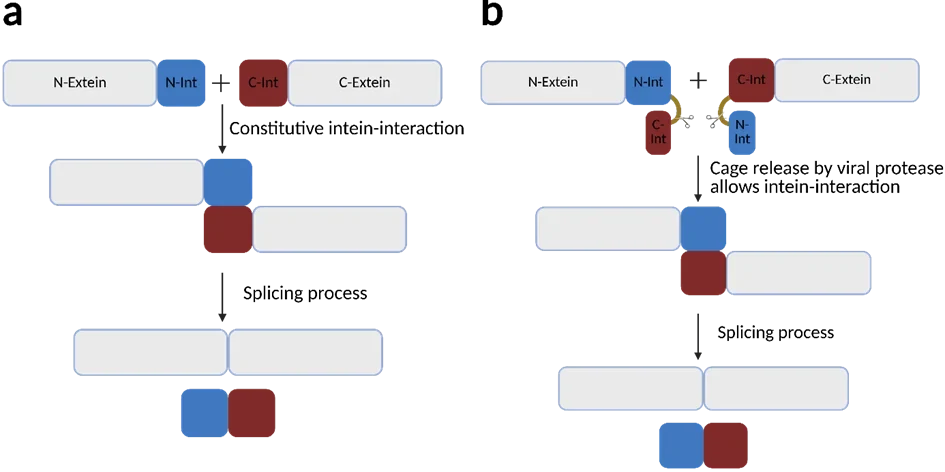
protein splicing. Splitted inteins domains (Npu from Nostoc punciforme) can excise themselves from a polypeptide construct and mediate the covalently relegation of the previously attached polypeptide (extein) (Figure 1a)(5,6). This can be further modified by caging the intein-fragments for controlling the timing of the splicing (Figure 1b). By integrating the potyvirus member specific cleavage site of the Nla-protease the splicing and therefore the relegation of the two extein-fragments is achieved only under the presence of the viral protease. (6)
Figure 1: Schematics of normal intein splicing (a) and modified caged intein splicing (b). In normal protein trans-splicing, the N-terminal (red) and the C-terminal (blue) intein-fragments interact with each other. Together they excise themselves out and religate their exteins together without a scar.5 To inhibit splicing activity, the intein fragments are caged with the capturing part of the other intein fragment without the catalysation domain. This blocks intein splicing until release of the cage via cleavage of the virus specific cleavage site, which is located along the protein linker (yellow).6 Created with Biorender.com.
For our project we are developing a tool-box, where an effector protein gets transferred into the nucleus upon abundance of the Nla-protease of a specific potyvirus. The tool consists of a caged intein construct, where the adjacent exteins are spliced together upon viral protease-based cage-release. The recompleted exteins are the N- and C-terminus of the plant’s nuclear localisation signal attached to a Cre recombinase. Therefore, upon infection the corresponding viral Nla-protease gets expressed and releases the intein-cage. The NLS of the Cre-recombinase gets completed via splicing, allowing for the import of the Cre-recombinase. The Cre recombinase is protein, which recognizes specific palindromic 34 bp. Sequences (LoxP sites) and based on two LoxP sites inverts or excises a in between laying sequence.8 The Cre recombinase elicits in our case the inversion of an inversed recombinant Barnase gene (strong unspecific RNase), inducing the unspecific shredding of RNA-transcript inside of the plant cell up to the cell death.9
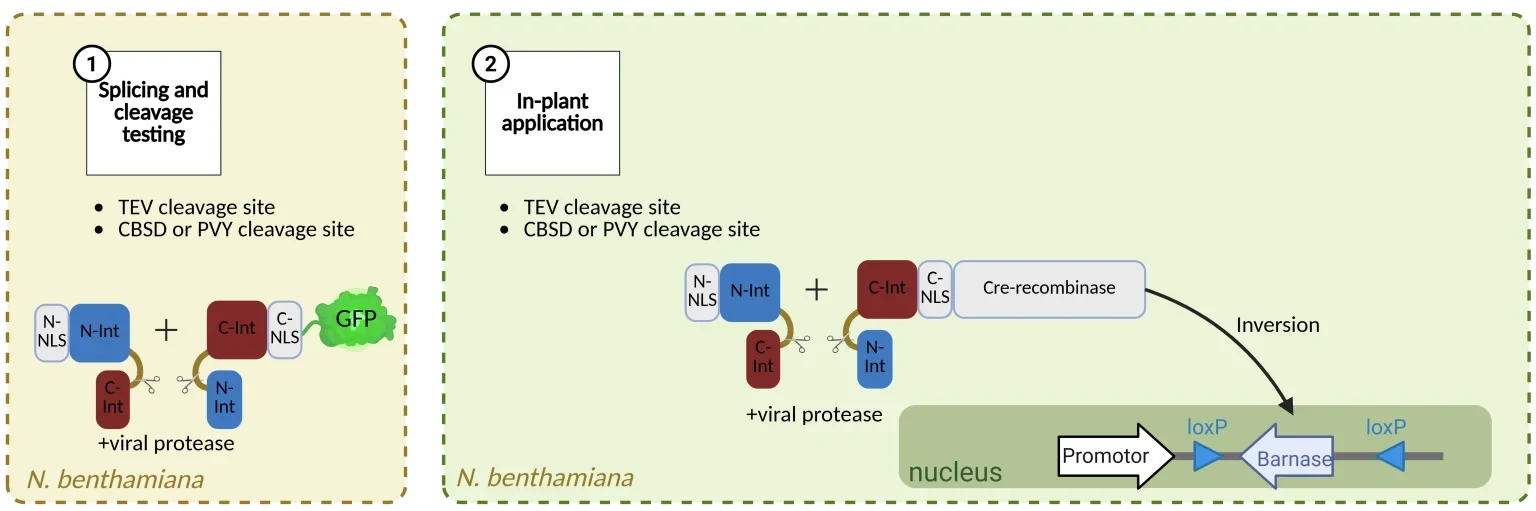
As the first step, we want to verify the splicing success of our construct. This is why we choose to firstly utilize the robust tabbacco etch virus proteases cleavage site (3). Initially we want to test it in Nicotiana benthamiana and with a GFP instead of the Cre Recombinase for easier visualization of the splicing success of our construct (Figure 2). Then we will complement this with the Nla-Pro cleavage sites for the cassava brown streak virus and for the potato virus Y (Figure 2).
Figure 2: Project draft for generating a resistance tool based on the potyviral protease activity. NLS nucleus localisation signal, Int intein-fragment (Red for C-terminal, Blue for N-terminal; Intein-fragments of the cage are incomplete fragments), GFP green fluorescent protein. CBSD for cassava brown streak virus protease, PVY for potato virus Y protease. Created with Biorender.com.
The advantage of our system lays in the transferability on all Nla-pro containing potyvirus members. Just the Cleavage site and the NLS sequence have to be adapted for the specific potyvirus species. This allows to generate plant resistance against a huge variety of potyvirus pathogens.
References
- Yang, X., Li, Y., & Wang, A. (2021). Research Advances in Potyviruses: From the Laboratory Bench to the Field. Annual Review of Phytopathology, 59(Volume 59, 2021), 1–29.
- Hýsková, V., Bělonožníková, K., Chmelík, J., Hoffmeisterová, H., Čeřovská, N., Moravec, T., & Ryšlavá, H. (2024). Potyviral Helper-Component Protease: Multifaced Functions and Interactions with Host Proteins. Plants, 13(9), 1236. mmm
- Xiao, H., Lord, E., & Sanfaçon, H. (n.d.). Proteolytic Processing of Plant Proteins by Potyvirus NIa Proteases. Journal of Virology, 96(2), e01444-21.
- Jia, Z., Rui, P., Fang, X., Han, K., Yu, T., Lu, Y., Zheng, H., Chen, J., Yan, F., & Wu, G. (2025). Proteolysis of host DEAD-box RNA helicase by potyviral proteases activates plant immunity. New Phytologist, 245(4), 1655–1672.
- Shepherd, C., Lawson-Williams, M., Holland, A., Bello, A. J., Sexton, D. W., & Olorunniji, F. J. (2025). Conditional Split Inteins: Adaptable Tools for Programming Protein Functions. International Journal of Molecular Sciences, 26(2), 586.
- Gramespacher, J. A., Stevens, A. J., Nguyen, D. P., Chin, J. W., & Muir, T. W. (2017). Intein Zymogens: Conditional Assembly and Splicing of Split Inteins via Targeted Proteolysis. Journal of the American Chemical Society, 139(24), 8074–8077.
- Tomlinson, K. R., Bailey, A. M., Alicai, T., Seal, S., & Foster, G. D. (2017). Cassava brown streak disease: Historical timeline, current knowledge and future prospects. Molecular Plant


